
So far I feel the reality is not living up to the hype or maybe I am penalised for not working with human/mouse/rat resequencing data. 
I am working under an ubuntu 64 bit environment and the data loading of one lane of Paired End reads was extremely slow.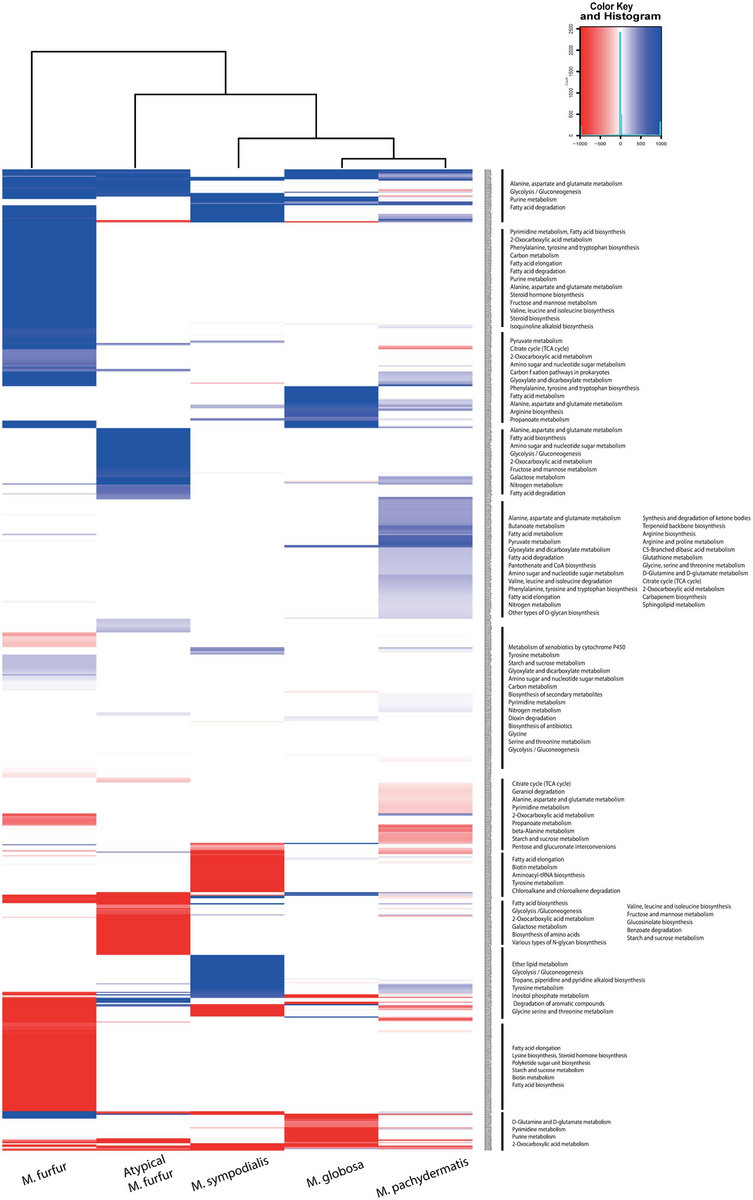 
I also find the comparison of their assembler with maq and soap a laugh, this is comparing an assembler with mappers. I do a lot of work with small ncRNAs and cannot find any tools in the trial that are remotely useful. The other question is - can we do real mapping with CLC or are we stuck with contig assembly (with or without reference). Does anyone have any clear instruction on sorting Illumina based indexed sequencing? I am just beginning to evaluate CLC Genomic Workbench for use with Illumina output and I am finding it so 454 orientated that it is driving me crazy with irrelevant instructions. Has anybody experience using their Genomics Server in combination with the workbench? It's supposed to let users run the workbench as a client and let the assembly and mapping to be calculated on the server, but again loading the results into the client for browsing could still be a bottleneck.įinally, what alternatives are there for browsing assembly/mapping results (when mapping to a reference genome) interactively and with some graphics, I mean for end users? I just read about MapView but haven't tested it yet. I guess they're still improving this kind of functionality. the assembled contigs table does not allow to search for gene names even though the reference is RefSeq mRNA from gene bank with lots of annotation. However, for the moment the search capabilities in the workbench is not yet as good as I'd like to have it, e.g. We're doing RNA-Seq (qualitative), and the main reasons why our biologists are interested in the workbench is to query for their favorite gene in the assembly and look how many reads align where - confirm the presence of transcripts and ultimately/hopefully work out tissue specific isoforms. Anyway, there's probably a trick to split thing up. Even if I did the assemblies and mappings for them, the resulting contig file is too large to load on any winXP machine (limited to <4 Gb of memory) for browsing. I run the workbench on a 64 Gb Linux machine, but our end users only have small winXP workstations. Still the application is memory greedy - the assembler/mapper seems to be a stand alone binary program (C/C++?) that's called by the workbench, whereas the rest is java which consumes lots of memory (~30 Gb when loading 7mio Solexa reads in fastq format and the human RefSeq mRNAs as the reference). ) and "only" consumes about 2Gb of memory. I like the user interface, and the assembly against a reference genome/transcriptome is fast (comparable with bowtie - not arguing about minutes. I am testing the CLC Genomic Workbench 3.5 for our molecular biologists (our main users). Īs Torst points out, the many helpful graphic utilities with CLC (presumably the reason it is slow?) make the experience more pleasant. on the other hand, CLC does give you the graphic that neatly lines up all the reads so you have the opportunity of looking at them to try to understand how it made its decisions. with CLC, staring at some of the contig ends after blasting them on what for sure is where they come together, and then looking at the read coverage in the unjoined region, it is hard to understand what kept the assemler from joining them into a larger contig. other problems, like accuracy, are introduced with reduced coverage_cutoff but at least it acts as one would expect. with velvet there is a large difference in the contig size when you reduce the coverage_cutoff. mismatch penalty of 2 gives basically the same result as mismatch penalty of 1 for de novo assembly. the penalty adjustments dont seem to do anything significant. my problem with CLC is related to connecting contigs that "by eye" have plenty of coverage at overlapping regions but CLC wont connect them. 
strangely enough I am now getting "contigs" in de novo assembly of 36 bases. One thing different in the new release is that there is no longer a minimum contig length of 200 bases.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
